New Chip Developed That Improves Testing and Tracing for COVID-19
0 View
Share this Video
- Publish Date:
- 2 May, 2021
- Category:
- Covid
- Video License
- Standard License
- Imported From:
- Youtube
Tags

Professor Jeremy Edwards of the University of New Mexico is researching new technology that can detect COVID and other respiratory viruses more quickly and accurately. Credit: University of New Mexico
Jeremy Edwards, director of the Computational Genomics and Technology (CGaT) Laboratory at the University of New Mexico, and his colleagues at Centrillion Technologies in Palo Alto, California and West Virginia University, have developed a chip that provides a simpler and faster method of genome sequencing for viruses such as COVID-19.
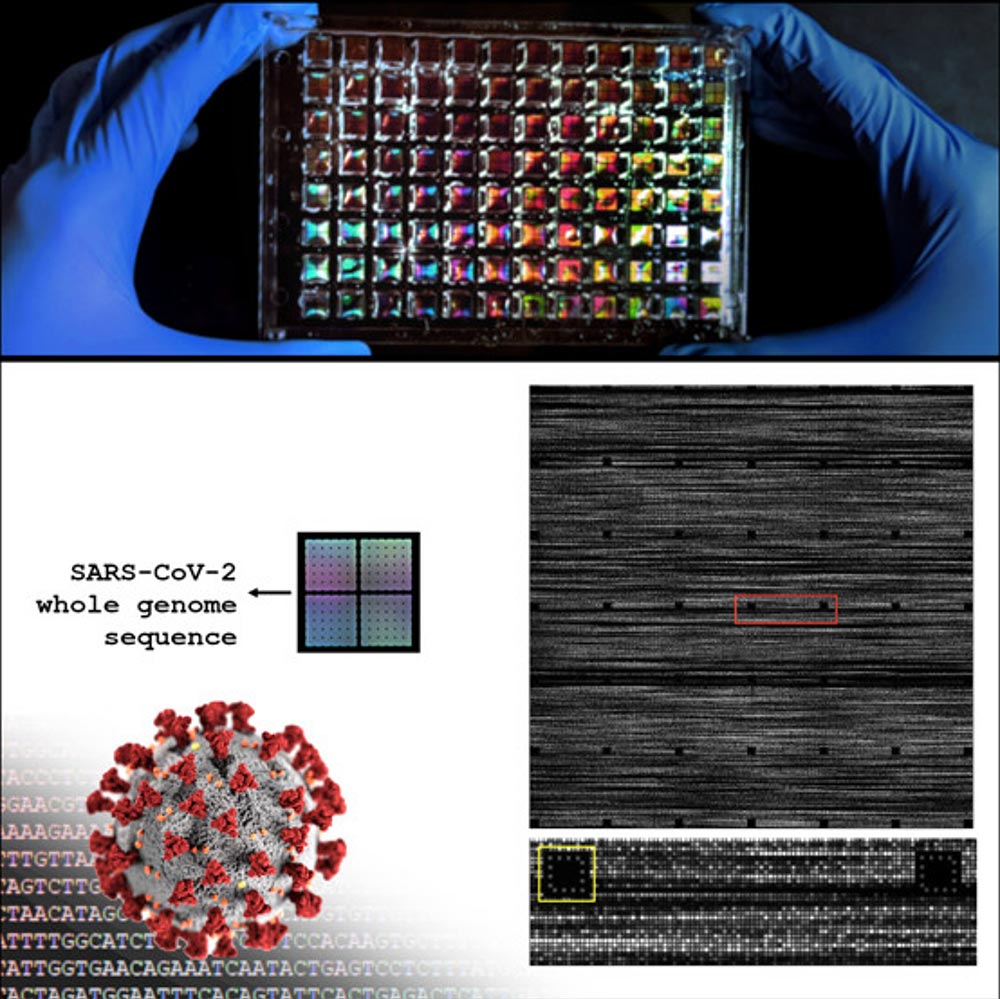
Their study, entitled “Highly Accurate Chip-Based Reclassification of SARS-CoV-2 Clinical Specimens” was recently published in Langmuir of the American Chemical Society. As part of the study, scientists created a tiled genome array that they developed for fast and inexpensive full viral genome resequencing and applied their SARS-CoV-2 specific genome tiling array to quickly and accurately rearrange the viral genome from eight clinical samples. obtained from patients. in Wyoming who tested positive for SARS-CoV-2. Ultimately, they were able to sequence 95 percent of each sample’s genome with an accuracy of more than 99.9 percent.
SARS-CoV-2 Complete Genome Sequence. Credit: University of New Mexico
“This new technology provides faster and more accurate tracking of COVID and other respiratory viruses, including the appearance of new variants,” said Edwards, a professor in the UNM’s Department of Chemistry and Chemical Biology. “With this simple and fast testing procedure, scientists will be able to monitor progress more closely and prevent the onset of the next pandemic.”
With more than 142 million people worldwide having contracted the virus, vigilant testing and contact tracking are the most effective ways to slow the spread of COVID-19. Traditional clinical testing methods often yield false positives or negatives, and traditional sequencing methods are time consuming and expensive. This new technology will virtually remove all these barriers.
“Since the document was filed, the technology has continued to evolve with improved accuracy and sensitivity,” said Edwards. “The chip technology is the best available technology for large-scale viral genome surveillance and monitoring of viral variants. This technology can not only help control this pandemic, but also prevent future pandemics. “
Reference: “High-precision chip-based reclassification of SARS-CoV-2 clinical specimens” by Kendall Hoff, Xun Ding, Lucas Carter, John Duque, Ju-Yu Lin, Samantha Dung, Priyanka Singh, Jiayi Sun, Filip Crnogorac, Radha Swaminathan , Emily N. Alden, Xuechen Zhu, Ryota Shimada, Marijan Posavi, Noah Hull, Darrell Dinwiddie, Adam M. Halasz, Glenn McGall, Wei Zhou, and Jeremy S. Edwards, April 13, 2021, Langmuir.
DOI: 10.1021 / acs.langmuir.0c02927
The mission of the Computational Genomics and Technology (CGaT) Laboratory is to provide training in bioinformatics research for undergraduate, graduate and Ph.D. students, as well as postgraduate fellows; provide collaborative research interactions to use bioinformatics computing tools for UNM researchers, and to conduct state-of-the-art and innovative bioinformatics and genomics research within the center.