COVID-19 Silences the Immune Response in Infected Cells in the Gut
0 View
Share this Video
- Publish Date:
- 27 April, 2021
- Category:
- Covid
- Video License
- Standard License
- Imported From:
- Youtube
Tags
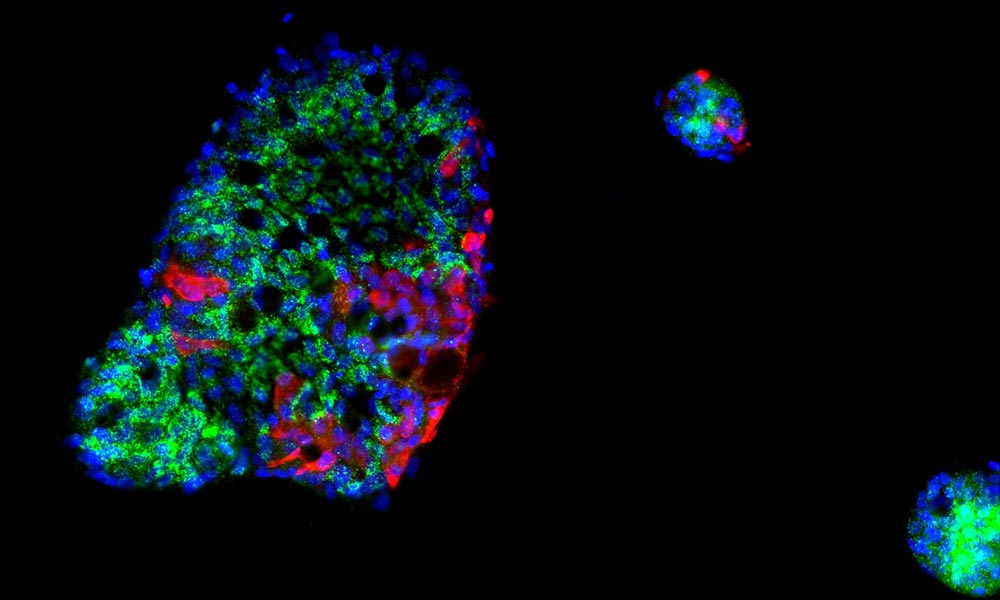
The researchers were able to monitor the growth of the virus in organoids derived from human intestinal cells. Pink and red show areas of SARS-CoV-2 infection. Credit: Mohammed Shahraz, Sergio Triana / EMBL; Camila Metz-Zumaran / University of Heidelberg
Scientists transform human gut cells into ‘mini intestines’ to monitor the infection process.
In an effort to determine the potential of COVID-19 to start in a person’s gut and to better understand how human cells respond to SARS-CoV-2, the scientists used human gut cells to make organoids – 3D tissue cultures derived from human cells, which mimic the tissue or organ from which the cells originate. Their conclusions, published in the journal Molecular Systems Biology, indicate the potential of infection to harbor in a host’s gut and reveal intricacies in the immune response to SARS-CoV-2.
“Previous research had shown that SARS-CoV-2 can infect the gut,” said Theodore Alexandrov, who leads one of the two EMBL groups involved. “However, it remained unclear how gut cells build their immune response to the infection.”
In fact, the researchers were able to determine the cell type most severely infected by the virus, how infected cells trigger an immune response, and – most interestingly – that SARS-CoV-2 silences the immune response in infected cells. These findings may shed light on the pathogenesis of SARS-CoV-2 infection in the gut, and indicate why the gut should be considered to fully understand how COVID-19 develops and spreads.
According to Sergio Triana, lead author and PhD student on EMBL’s Alexandrov team, the researchers observed how infected cells appear to start a cascade of events that produce a signaling molecule called interferon.
“Interestingly, although most of the cells in our mini-gut had a strong immune response elicited by interferon, SARS-CoV-2 infected cells did not respond in the same way and instead showed a strong pro-inflammatory response”, Sergio says. “This suggests that SARS-CoV-2 interferes with the host’s signals to disrupt an immune response at the cellular level.”
Coronaviruses, including SARS-CoV-2, cause infection by adhering to specific protein receptors found on the surface of certain cell types. Among these receptors is the protein ACE2. Interestingly, the researchers showed that the infection is not explained solely by the presence of ACE2 on the surface of the cells, underscoring our still limited knowledge about COVID-19, even after a year of massive research efforts worldwide.
As the disease progressed in the organoids, the researchers used single-cell RNA sequencing, which involves a variety of techniques to amplify and detect RNA. Of these single-cell technologies, Targeted Perturb-seq (TAP-seq) provided sensitive detection of SARS-CoV-2 in infected organoids. Lars Steinmetz’s research group at EMBL recently developed TAP-seq, which the researchers combined with powerful computing tools, enabling them to detect, quantify and compare the expression of thousands of genes in individual cells in the organoids.
“This finding could provide insight into how SARS-CoV-2 protects itself from the immune system and offers alternative ways to treat it,” says Lars. “Further research can help us understand how the virus grows and the different ways it affects the human immune system.”
Reference: April 27, 2021, Molecular Systems Biology.
DOI: 10.15252 / msb.202110232